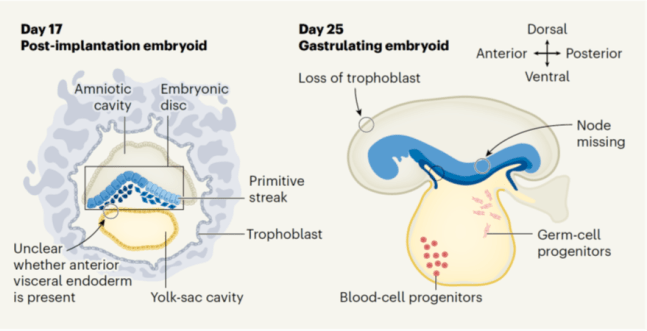
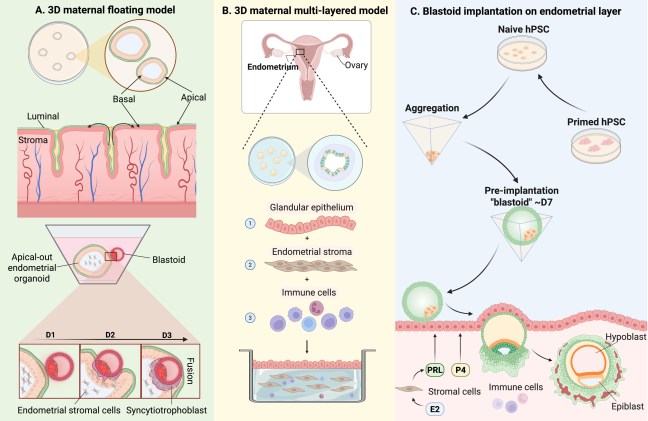
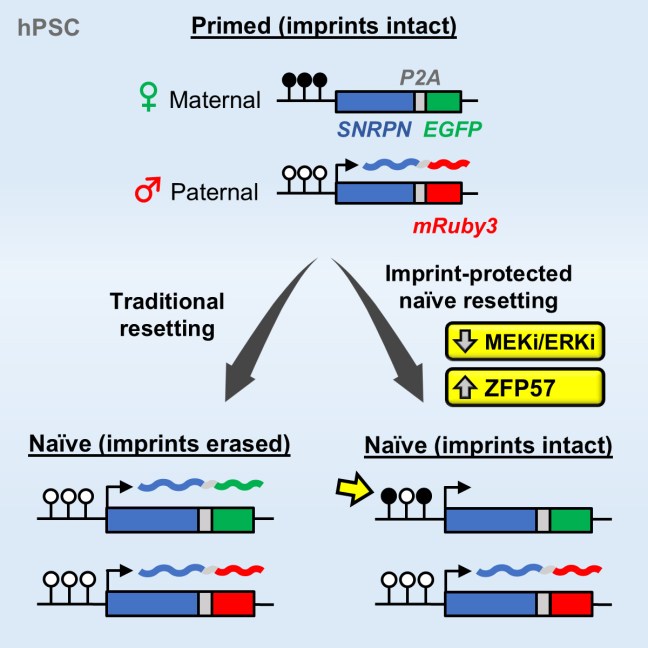
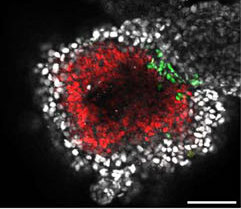
Our study on tracking and mitigating imprint erasure during induction of naive human pluripotency is now online at Stem Cell Reports. In this paper, we tackled a long-standing problem in stem cell biology: the erosion of parent-specific epigenetic marks (so-called “imprints”) during the generation of naive stem cells, which more closely resemble the pre-implantation blastocyst. These imprints are protected during the genome-wide wave of demethylation in the pre-implantation embryo and ensure monoallelic expression of hundreds of genes during human development. However, imprints are demethylated under current conditions for inducing naive human pluripotency, which creates an epigenetic imbalance in naive stem cells and their derivatives. To track the dynamics of imprint erasure during naive resetting in real time, we established a dual-colored fluorescent reporter at both alleles of the imprinted SNRPN locus. During primed-to-naive resetting, SNRPN expression becomes biallelic in most naive cells, and biallelic SNRPN expression is irreversible upon re-priming. We utilized this live-cell reporter to evaluate chemical and genetic strategies to minimize imprint erasure. Decreasing the level of MEK/ERK inhibition or overexpressing the KRAB zinc-finger protein ZFP57 protected a subset of imprints during naive resetting. Combining these two strategies protected imprint levels to a further extent than either strategy alone. This study offers an experimental tool to track and enhance imprint stability during transitions between human pluripotent states in vitro.
This study was led by our former graduate student and postdoc, Laura Fischer, with support from our collaborators in the research groups of Sabine Dietmann and Ting Wang at WashU and Harald Jueppner at MGH.